Molecular chaperones
【What are molecular chaperones?】 Molecular chaperones are originally defined as a protein family that assists in protein folding. Now, they are known as a supporting role with a strong presence that supports not only folding but also the “life of the protein” in the cell.
【Anfinsen’s dogma】In the 1950s, Christian Anfinsen and his colleagues demonstrated that protein folding is a spontaneous process. There are several ways to express it.
-Amino acid sequence of the protein dictates the structure.
-Protein folds into a native structure that has the minimum Gibbs free energy.
These statements are a fundamental principle of protein folding. Since the amino acid sequences are encoded in genomes, and are translated to produce polypeptide chains at the ribosome via the mRNA, the protein folding is a simple process in the final step of the Central Dogma of Biology (DNA -> RNA -> protein).
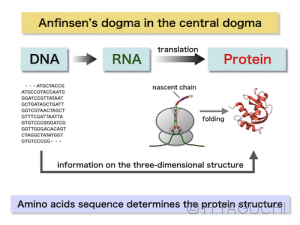
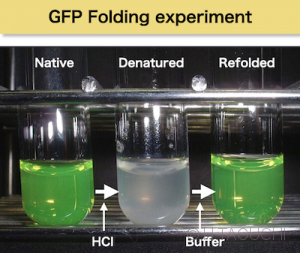
Figure below shows a demonstration experiment to visualize spontaneous folding using Green Fluorescent Protein (GFP). GFP has a beautiful green fluorescent, the color of GFP is dependent on the correct tertiary structure. The green color is instantly disappeared after GFP is denatured by adding HCl to lower the pH (the slightly cloudy material is due to aggregation after denaturation). Then, after the pH of the solution is neutralized by adding a buffer, the green color gradually returns.
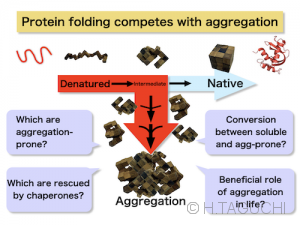
【Folding vs. Protein aggregation】 Protein folding always competes with aggregate formation. Typical globular protein usually has a hydrophobic core inside the native structure. Hydrophobic amino acids to form the core are exposed after the protein denaturation, or during the protein synthesis at the ribosome. Since hydrophobic molecules tend to associate with each other in aqueous solutions (hydrophobic interaction), the intramolecular hydrophobic interaction drives the correct protein folding. However, intermolecular hydrophobic interaction between denatured proteins leads to form aggregates. Typically, aggregate formation is a higher-order reaction, leading to the irreversible aggregates. The most common protein aggregates in our daily life would be boiled eggs.
The aggregate formation is dependent on protein concentration. It is known that the inside of a cell is very crowded with proteins and other biological macromolecules. This is not suitable environment for protein folding.
【Molecular chaperone】In the crowded cellular environment, there are proteins that help protein folding to prevent irreversible aggregate formation. This is the molecular chaperone. The concept of chaperones was first proposed in the late 1980s, when the intracellular functions of major chaperones such as chaperonins (GroEL, Hsp60, CCT, etc.) and Hsp70/DnaK became known. The original definition of a molecular chaperone by John Ellis was “a protein that helps another protein fold its correct structure but does not become its own final component”.
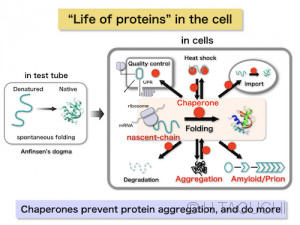
【Concept of Life of proteins, Proteostasis】The concept of chaperones originated from their ability to assist in the protein folding, but it has since become clear that chaperones are involved in a wide variety of events involving proteins in the cell. If we think of the “birth” of a protein as the point where it is synthesized in the ribosome, it “grows up” to a correct structure of the protein. Protein in the cell faces to “stresses” such as heat shock, moves to a different location, forms abnormal structures such as amyloids, and finally dies….. Thus, proteins in the cell are associated with many kinds of chaperones from the cradle to the grave. Alternatively, chaperones are essential to maintain protein homeostasis (proteostasis) in the cell. In order to understand proteins in the cell, it is essential to understand chaperones.
【Selected publications】
-Miwa T, Chadani Y, *Taguchi, H.Escherichia coli small heat shock protein IbpA is an aggregation-sensor that self-regulates its own expression at post-transcriptional levels.
Mol Microbiol 115:142-156 (2020) doi: 10.1111/mmi.14606.
-Okuda, M, Niwa, T., *Taguchi, H.,
Single-Molecule Analyses on the Dynamics of Heat Shock Protein 104 (Hsp104) and Protein Aggregates.
J. Biol. Chem. 290, 7833-7840 (2015)
-Niwa, T., Kanamori T., Ueda, T.*, Taguchi, H.*
Global Analysis of Chaperone Effects Using a Reconstituted Cell-Free Translation System
Proc. Natl. Acad. Sci. U.S.A. 109, 8937-8942 (2012)
-Fujiwara, K., Ishihama, Y., Nakahigashi, K., Soga, T. and Taguchi, H.*
A systematic survey of in vivo obligate chaperonin-dependent substrates.
EMBO J. 29, 1552-1564 (2010)
-Niwa, T., Ying, B.-W., Saito, K., Jin, W. Z., Takada, S., Ueda, T.*, Taguchi, H.*
Bimodal protein solubility distribution revealed by an aggregation analysis of the entire ensemble of Escherichia coli proteins.
Proc. Natl. Acad. Sci. U.S.A. 106, 4201-4206 (2009)